


Artificial intelligence-driven platform identifies new ALS gene targets
Twenty-eight targets were identified, and the collaborative study was led by Insilico with support from Answer ALS and researchers at nine institutions including the Johns Hopkins University School of Medicine
Amyotrophic Lateral Sclerosis (ALS) affects 5 to 10 out of every 100,000 people. Despite being known to medicine for nearly two centuries, few treatments exist, and none can alter the course of disease. More than 90% of ALS cases are sporadic, with no known family history or cause of illness. With a wide heterogeneity in the age of onset and clinical course, scientists have struggled to identify the molecular causes of ALS that can lead to therapies.
To bridge this gap, scientists launched the Answer ALS initiative five years ago to collect cells and DNA from 1000 ALS patients. The goal was ambitious: to create nervous system cell lines from each patient for detailed molecular analysis to identify subtypes of disease, with the hopes that these would lead to targeted therapies. The cell lines serve as nervous system biopsies, like those used in cancer, to provide enormous amounts of individual patients biological “fingerprints”. In a new paper in Frontiers in Aging Neuroscience, an international team of scientists, including Jeffrey Rothstein MD, PhD, Founder and Director of the Packard Center, used Answer ALS data to identify novel drug targets using an artificial intelligence platform from Insilico Medicine called PandaOmics.
“We are truly excited to see the Answer ALS data being used to identify possible ALS disease-causing pathways and candidate drugs,” said Jeffrey D. Rothstein MD, PhD, Founder and Director, Robert Packard Center for ALS Research and Answer ALS. “The work by Insilico is exactly how this unprecedented program was envisioned to help change the course of ALS.”
“I think AI is the future because it can interpret huge amounts of data and generate novel hypotheses,” says Frank Pun, head of Insilico‘s Hong Kong team and first author of the new paper.
Since the first ALS-linked gene was discovered in 1993, researchers have discovered over 50 genes linked to the condition. Advances in molecular biology mean that researchers know more about the cellular factors that contribute to disease than ever before. This knowledge, however, hasn’t led to effective treatments. But these genetic changes do not necessarily explain the molecular defect that underlies that vast majority of ALS patient with sporadic, not familial, ALS. What’s more, researchers have uncovered substantial heterogeneity among people with sporadic ALS with regards to age of onset, rate of disease progression, and more. The biological information, gathered for the Answer ALS cell lines provide researchers with possible tools to uncover patient subgroups and candidate druggable pathways.
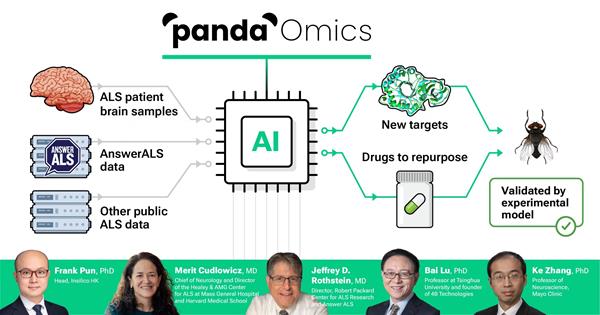
Pun and colleagues used PandaOmics to analyze the RNA and proteins (known as transcriptomics and proteomics) of post-mortem neural tissue and neurons derived from induced pluripotent stem cells from ALS patients and healthy controls, including those generated as part of Answer ALS. PandaOmics’s analysis of the transcriptomic and proteomic data used machine learning to predict the genes that might be associated with ALS. The software platform can also analyze how well a particular gene might be suited for the development of potential therapies.
This first run yielded a list of 28 potential candidates: 17 high-confidence and 11 novel therapeutic gene targets. To validate that the promising targets impacted ALS, the researchers turned to a Drosophila fruit fly model of ALS caused by a repeat expansion in the C9orf72 gene. The researchers screened 26 of the 28 targets identified by the PandaOmics analysis in the fruit fly model (2 targets lacked Drosophila equivalents). The researchers suppressed expression of these genes with RNA interference (RNAi) and measured whether this rescued neurodegeneration in the C9 fruit fly eye. Eight of the novel targets strongly showed this effect.
Pun says that these results help to validate the strength of the PandaOmics platform in its ability to identify actionable therapeutic gene targets of interest. Further assays need to be performed to show exactly how these targets might be involved in ALS, and how they might be targeted by small molecules, he concludes.
